
Introduction
Computational Biology employs powerful tools from computer science, applied mathematics, and statistics to solve a diverse array of biological problems. Major avenues of research include genome assembly--piecing together a large collection of short DNA sequences, gene finding--locating patches of DNA that have biological function, genomic sequence alignment--the ordering of multiple sequences to elucidate their parallel structure, protein structure prediction--determining the 3D structure of proteins from their chemical makeup, and phylogenetic analysis--studying and modeling evolutionary relationships between species. Mathematical and computational techniques are continuously being developed, analyzed, and improved to offer greater utility in biological arenas. A solid grounding in this rapidly advancing field requires an understanding of both the computational/mathematical aspects as well as the underlying biology of this interdisciplinary field. In this preliminary chapter, we discuss basic molecular biology as well as a popular application to illustrate the usefullness of computational techniques in biology.
Molecular Biology
The Cell

The cell is the fundamental working unit of living organisms. Some organisms are unicellular and have only a single cell (e.g. bacteria), whereas other organisms are multicellular and can have trillions (e.g. humans). Cells contain "organelles"--subcellular differentiated structures that perform specific functions--that work together to accomplish tasks needed for maintenance and proliferation. The most major of these is the nucleus, the organelle in eukaryotic ("with nucleus") cells that houses the cell's chromosomes and thereby genetic material in the form of DNA. The nucleus sits in the cytoplasm, the jelly-like substance that houses the organelles. Not all cells have nuclei though--some cells are prokaryotic ("without nucleus") and their DNA simply resides in the cytoplasm. There are many other important organelles in the cell, the most major of which include:
- Cell(Plasma) Membrane: The outer membrane of the cell that surrounds the cytoplasm and separates the cell from its extracellular environment.
- Ribosomes: Complexes composed of ribosomal RNA and proteins responsible for converting DNA's transcribed message into proteins/protein subcomplexes (polypeptides).
- Mitochondria: Organelles whose primary responsibility is to convert organic materials into energy that the cell can use.
DNA and RNA

DNA
Deoxyribonucleic acid (DNA) is a nucleic acid that usually appears in the form of a double helix and serves as the primary and permanent storage of information for the organism. DNA is a long polymer of nucleotides ("bases") and segments of DNA called genes encode sequences of amino acids, the building blocks of proteins. Variations in DNA manifest into an organism's unique phenotype (its observable biochemical and physical characteristics) through biology's actors, proteins. In DNA, nucleotides are composed of (i) a phosphate group, (ii) a deoxyribose sugar, and (iii) one of four nitrogen-containing bases: adenine(A), guanine(G), cytosine(C), and thymine(T). Each strand of the double helix consists of a long sequence of nucleotides covalently linked to each other. The two strands themselves bond to each other with hydrogen-bonds and in a complementary fashion: A is complementary to T (double H bond), G is complementary to C (triple H bond). Because of this complementarity, one strand can be inferred entirely from its complementary sequence and so each of the two strands contains the same amount of information as the two strands together.
Because of the asymmetric linkage of nucleotides, a strand of DNA has an inherent directionality. That is, each of the two strands of the double helix has an orientation. Using chemical nomenclature, one direction of a single strand is called the 5' direction and the other is the 3' direction. The two strands of the double helix bond to each other in such a way that the 3' direction of one strand pairs with the 5' direction of the other strand. This property of DNA makes it antiparallel.
RNA
Ribonucleic acid (RNA) is a nucleic acid that can appear in a diverse variety of forms and shapes. Unlike DNA, RNA is almost always single-stranded. It is also much shorter and contains ribose as its sugar as opposed to deoxyribose. Finally, RNA uses the base uracil(U) instead of DNA's thymine(T).
Proteins

Proteins are macromolecules that carry out the tasks of the cell and are essential to all living cells. They are organic compounds composed of amino acids which are joined to one another by peptide bonds. Proteins undertake structural, mechanical, enzymatic, signalling, and regulatory functions in organisms.
Amino Acids
Amino acids, the building blocks of proteins, connect to each other with peptide bonds to form proteins/protein subcomplexes. There are 20 amino acids and each has the same general structure but differ in their unique side chains. Side chains cause different amino acids to have different biochemical properties. The sequence of amino acids is known as the primary structure of a protein.
Higher Structure
Proteins fold into complex shapes to allow them to carry out their function. They fold into unique 3-dimensional structures that scientists characterize with three higher levels of structure, along with the primary structure (just the sequence):
- Secondary Structure: Commonly-observed sub-structures of the protein that includes alpha helices and beta sheets.
- Tertiary Structure: The collective shape of a single folded protein. That is, the overall combination of secondary structures and their interaction with each other.
- Quartenary Structure: Structure composed of the joining of multiple protein subunits.
The Genome

All of the genetic information of an individual is stored in his or her DNA. DNA coils and compacts tightly into chromosomes. Humans have 23 pairs of chromosomes and the full DNA sequence of an organism is called its genome. Each cell in an organism that contains the complete copy of the genome contains all of its genetic information.
Chromosomes can be subdivided into genes, patches of DNA that encode proteins. Within these genes, there are portions known as exons and portions known as introns. As described below, exons are sequences that end up being expressed into protein (via the intermediate mRNA), and introns are spliced out. Upstream (in the 3' direction) of the gene is the promoter, a regulatory sequence that interacts with proteins called transcription factors to control the transcriptional activity of the gene. The DNA sequences between genes are known as intergenic sequences.
The Standard Flow of Information
The central dogma of molecular biology deals with the flow of information from nucleic acid to protein. It states that once genetic information has reached the protein form, it cannot be transferred from protein to either protein or nucleic acid. It is often confused with the standard flow of information of the cell: DNA is converted into RNA which is then converted into protein. This standard flow is a cornerstone of molecular biology and its importance to our understanding of life cannot be underestimated.
Replication
Replication is the process by which the cell makes a copy of its own DNA. Each parental strand of the double helix serves as a template and replication relies on A-T and G-C complementary base-pairing. There is a fixed replication error rate of 1 in 10^9 to 10^10 base pairs, and this fixed error rate is the source of single nucleotide polymorphisms (SNPs), which are variations that occur in DNA when a single nucleotide is alterred.
The cell replicates its DNA in preparation for cellular division through one of two processes: mitosis or meiosis. During mitosis, a cell with a copied genome divides once and thereby forms 2 daughter cells, each containing a complete chromosomal complement. Cells undergo mitosis to generate identical cells to the parent cell. During the process of meiosis, a cell with a copied genome undergoes an initial cellular division and those two cells then undergo another division. Through the molecular mechanism of recombination, meiosis results in 4 daughter cells that are NOT identical to the parent cell, and contain only half of the full chromosomal complement (each of the 4 has 23 single chromosomes, instead of 23 pairs). Meiosis is used to generate sperm and eggs for the organism.
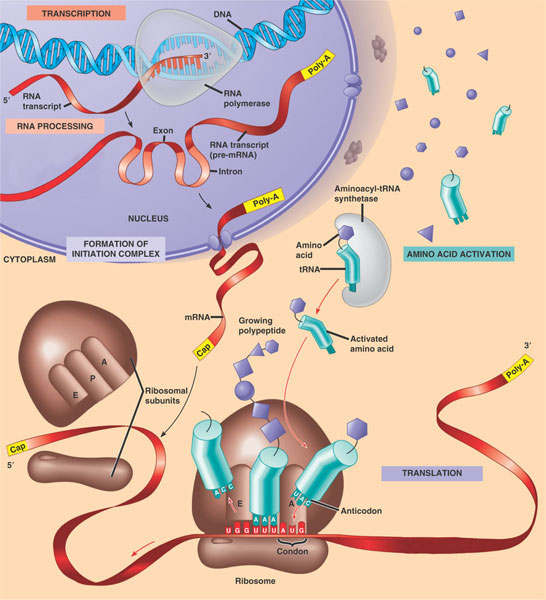
Transcription
Transcription is the process in which the cell synthesizes mRNA (messenger RNA) from a portion of DNA corresponding to a gene. DNA is transcribed to RNA through enzymatic action of RNA polymerase and the transcribed RNA is complementary to its template DNA. Due to this complementarity, the RNA contains all of the information of the portion of the DNA that was transcribed. This single-sranded RNA is referred to as mRNA for messenger RNA, since it will be carrying the message stored in the DNA. After being copied, the mRNA's message can be modified through the process of splicing: removing selective portions of the mRNA. Additionally, a poly-A tail is added to the 3' end of the mRNA along with a 5' cap--both of which offer stability to the mature messenger mRNA. In eukaryotes, everything up to this point has occured in the nucleus. Mature mRNA then leaves the nucleus and enters the cytoplasm to begin the next process, translation.
Translation
Translation is the process in which ribosomes in the cytoplasm guide the assembly of a sequence of amino acids based precisely on a sequence of mRNA. This sequence of amino acids is called a polypeptide and is specified through the order and content of bases of the mRNA. Another type of RNA, transfer RNA (tRNA) is also a major player in this process. tRNA functions to bring the individual amino acids corresponding to the sequence of the mRNA, to the ribosomes.
An Application of Computational Biology: Microarray Analysis

One of the most widely researched and utilized applications of computational biology is microarray analysis. Since the first development of cDNA microarrays by Pat Brown of Stanford University in 1995, a wealth of research has been devoted to finding the best ways to design the chips themselves and subsequently interpret the incredible amount of data that they produce. Microarrays can report the mRNA levels of many thousands of genes simultaneously and effectively. It is believed that the many genes of an organism's genome interact with each other in complex ways to regulate the actions of the cell. Microarray analysis provides a way to profile the transcriptional status of the cell--the amount of every gene's mRNA--and compare transcriptional responses to different stimuli. Moreover, algorithms and techniques have been developed to concisely interpret, visualize, and model these responses in such a way as to elucidate and concisely summarize their biological relevance. Microarray analysis depends heavily on the tools and techniques of computational biology, and without them, it would not be possible to process and interpret the large sets of multidimensional data that microarrays produce. Computational biology has provided invaluable techniques and tools that are no short of mandatory for successful interpretation and use of microarray technology.
Specific Application: Cluster Analysis 
Microarrays reveal the mRNA levels of genes via a scanned image. The image captures, for example, fluorescence for each gene that corresponds to the amount of mRNA present in the sample. These quantifications of the fluorescence have to first be normalized and corrected to adjust for systematic variation and other error, and then analysis can be performed. One of these forms of subsequent analysis is cluster analysis. Cluster analysis aims to divide similar genes into groups designed to provide insight into clusters of co-regulated genes. These partitions can then be used for a variety of purposes including finding regulatory mechanisms that control a group of coregulated genes, finding common sequences (motifs) in a group of coregulated genes, and discovering the underlying biological symmetries of the experiment at hand.
To accomplish this grouping task, we require the ability to store massive amounts of biological data, as well as the ability to process and perform intense computations with these data. On the design side, mathematically and computationally-efficient methods have been developed and programmed to process these data, incorporating a host of statistics and informatics. Each gene can be represented as a data point in n-dimensional real space with its coordinates representing its mRNA amounts as determined by the microarrays. Distances and correlations can be obtained between points and sets and, algorithmically, points can be allocated to clusters, sent from one cluster to another, or excluded all together. All of these tasks require intense computational capabilities integrated with a solidly laid foundation of mathematics and computer science and consequently, this is precisely a task for Computational Biology.

Comments (0)
You don't have permission to comment on this page.